How To Calculate Cdna Concentration From Rna

Ever wondered what makes that tiny sample of genetic material tick? Well, today we're diving into a surprisingly fun and incredibly useful topic: figuring out how much cDNA you have, starting from your RNA. Now, before you picture complicated lab coats and bubbling beakers, relax! This is all about understanding the basics, and it's more accessible than you might think.
Why is this even a thing? Think of RNA as a temporary messenger from your DNA, carrying instructions. To work with these instructions, especially for things like detecting specific genes or building DNA libraries, scientists often convert RNA into cDNA (complementary DNA). This cDNA is more stable and easier to manipulate. For beginners in the world of molecular biology, understanding cDNA concentration is like learning to read the ingredients list on a recipe – essential for success!
For families or hobbyists interested in science projects, this might sound a bit advanced, but imagine this: you're curious about what's happening in a plant's leaves or what makes certain yeast strains behave differently. If you're working with kits that involve DNA analysis or gene expression studies (yes, these exist for educational purposes!), knowing your cDNA concentration is key to getting reliable results. It's like measuring the flour accurately before baking your cake – you need the right amount for it to turn out!
Must Read
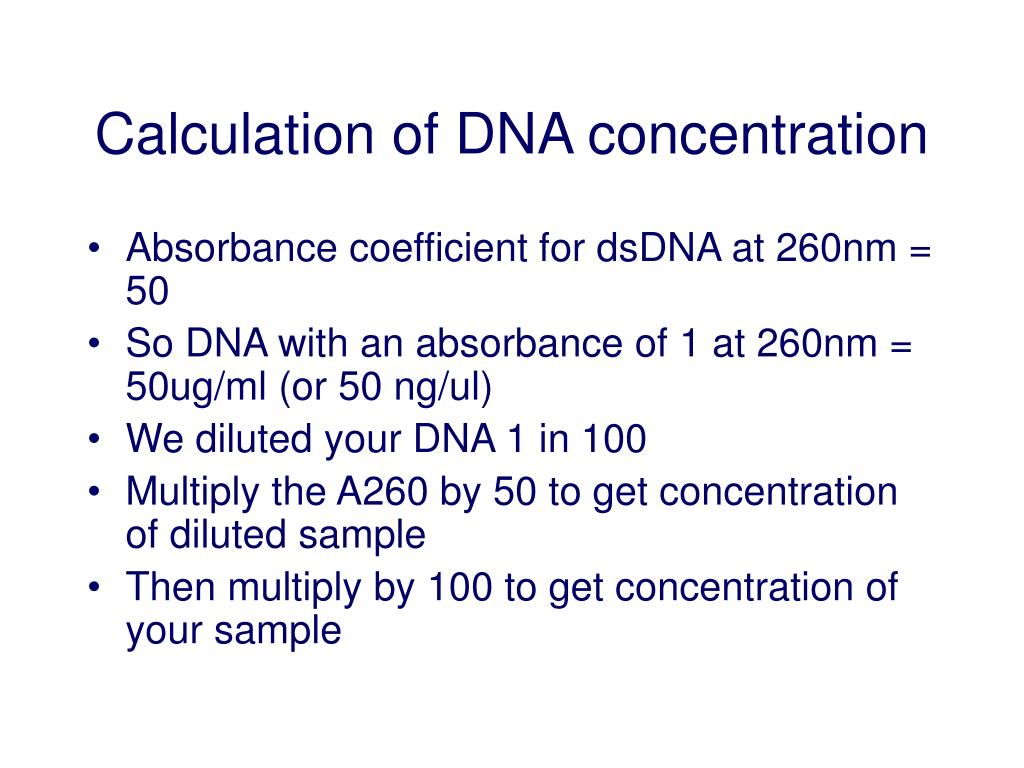
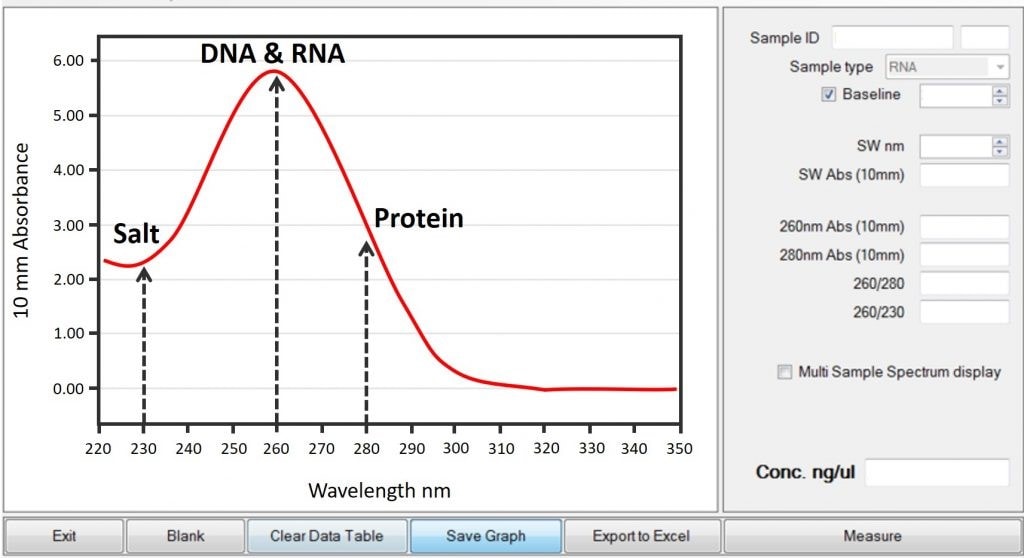
So, how do we actually get this number? The most common way is using a spectrophotometer. This nifty device measures how much light a sample absorbs at specific wavelengths. For nucleic acids like RNA and cDNA, we're usually interested in two wavelengths: 260 nanometers (which is where these molecules absorb light best) and 280 nanometers (which helps us detect protein contamination). The ratio of absorbance at 260nm to 280nm (the A260/A280 ratio) gives us an idea of the purity of our sample. A ratio between 1.8 and 2.0 is generally considered good for DNA and cDNA.
The actual concentration calculation is pretty straightforward once you have your absorbance readings. Most spectrophotometers will even do the math for you! Generally, the formula is: Concentration = Absorbance at 260nm × Dilution Factor × Constant. The constant is a pre-determined factor that converts absorbance units into concentration units (like nanograms per microliter, or ng/µL). If you're using a kit, it will often provide the specific constant to use.

Want to dip your toes in? If you're looking at DIY science kits that involve nucleic acid work, pay close attention to the instructions on sample preparation and measurement. Many kits might use simpler methods, like colorimetric assays, which also rely on color changes to indicate concentration. The principle is the same: measure something to figure out how much of your target molecule you have.
Don't be intimidated! Start by understanding the core concept: we're using light absorption to tell us about the quantity and quality of our genetic material. It's a fundamental skill that opens doors to so many exciting biological investigations. So, the next time you hear about cDNA concentration, remember it's not some arcane secret, but a practical step in unlocking the secrets of life, one measurement at a time. It’s a small step that leads to big discoveries, and that’s pretty rewarding!